Welcome to micapipe’s documentation! v0.2.3






Micapipe is a processing pipeline providing a robust framework to analyze multimodal MRI data. This pipeline integrates processing streams for T1-weighted, microstructure-sensitive, diffusion-weighted, and resting-state functional imaging to facilitate the development of multiscale models of neural organization. For this purpose, we leverage several specialized software packages to bring BIDS-formatted raw MRI data to fully-processed surface-based feature matrices.
Breaking news ️📰
micapipe version 0.2.3 Northern Flicker is finally here! Optimized to process 3T and 7T datasets!
About 👁️🗨️
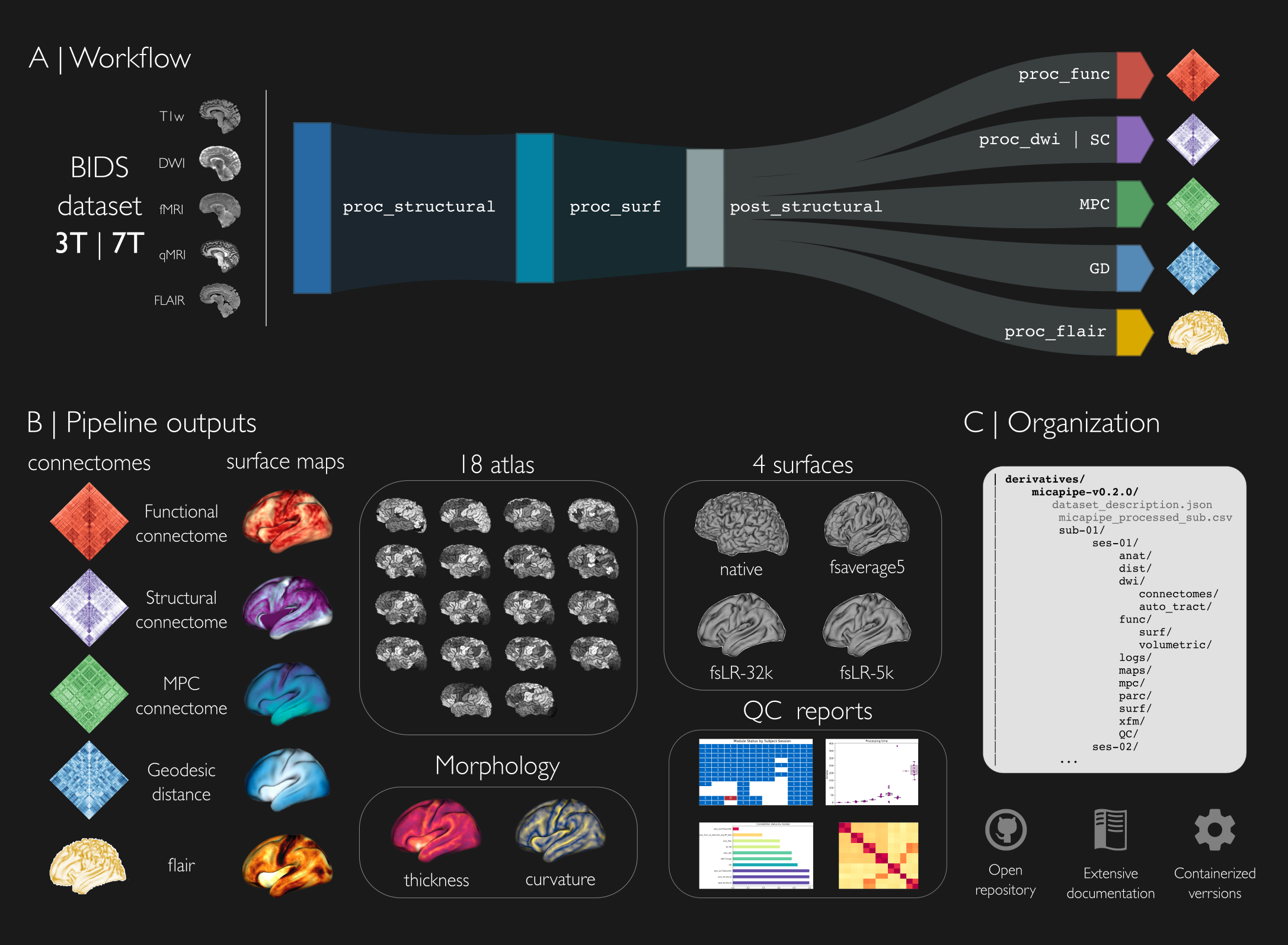
Micapipe generates systematic descriptions of cortico-cortical microstructural similarity, functional connectivity, structural connectivity, and spatial proximity. We hope that this open tool will be of use to researchers studying human brain structure and function across different spatial scales. The connectomes can be generated across 18 different cortical parcellations (100-1000 parcels), in addition to subcortical and cerebellar parcellations. All results are mapped to three different surfaces spaces: native, conte69 and fsaverage5, and all outputs are hierarchically ordered with BIDS conformed naming.
Reproducibility 👯♀️
To encourage reproducibility and robustness of investigations using micapipe, we provide a fully containerized version of the pipeline in the form of a Docker container. Step-by-step tutorials are provided for bare metal and containerized installations. We encourage users to use containerized versions, offered through Docker and Singularity, given the large number of software dependencies used by the pipeline to handle multiple MRI data modalities.
Datasets 🕵️♀️
Micapipe has been tested on several locally acquired datasets, as well as openly available repositories such as Microstructure-Informed Connectomis (MICA-MICs) Cambridge Centre for Ageing and Neuroscience (Cam-CAN), EpiC-UNAM, Midnight Scan Club, Auditory localization with 7T fMRI, SUDMEX_CONN and HCP.
Development and getting involved 🙋♀️
Should you have any problems, questions, or suggestions about micapipe, please post an issue or formulate a pull request on our repository.
Core development team 🧠
Micapipe is developed by members of the MICA-lab (https://mica-mni.github.io) and collaborators at the McConnell Brain Imaging Centre of the Montreal Neurological Institute.
Raúl Rodríguez-Cruces, MICA Lab - Montreal Neurological Institute
Alex Ngo, MICA Lab - Montreal Neurological Institute
Jessica Royer, MICA Lab - Montreal Neurological Institute
Peer Herholz, NeuroDataScience, ORIGAMI lab - Montreal Neurological Institute
Jordan DeKraker, MICA Lab - Montreal Neurological Institute
Yougeun Hwang, MICA Lab - Montreal Neurological Institute
Donna Cabalo, MICA Lab - Montreal Neurological Institute
Nicole Eichert, Jesus College, Oxford
Oualid Benkarim, MICA Lab - Montreal Neurological Institute
Yezhou Wang, MICA Lab - Montreal Neurological Institute
Sara Larivière, MICA Lab - Montreal Neurological Institute
Casey Paquola, MICA Lab - Montreal Neurological Institute
Boris Bernhardt, MICA Lab - Montreal Neurological Institute
